Transcriptomics-Driven Biomarkers for Major Depressive Disorder (MDD)
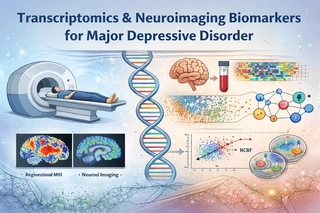
Project Overview
Major Depressive Disorder (MDD) is a severe psychiatric condition with complex functional and molecular mechanisms. Traditional structural imaging biomarkers have limited reproducibility and sensitivity — functional biomarkers like regional homogeneity (ReHo) demonstrate much stronger effects in large cohort studies.
This project adopts a transcriptomics-driven multi-modal framework to link RNA-seq–derived gene expression signatures to functional neuroimaging patterns in MDD, providing mechanistic insights and high-value biological interpretation for computational psychiatry.
Publication Reference:
Multimodal Neuroimaging and Transcriptomic Correlates of Major Depressive Disorder – JAMA Psychiatry
🔑 Key Scientific Insights
Functional brain alterations outperform structural markers:
ReHo-based functional MRI deficits show 2–3× larger effect sizes than cortical thickness reductions, confirming that MDD is primarily a disorder of functional dysregulation rather than structural atrophy.Robust hypoperfusion patterns:
ReHo deficits strongly co-localize with regionally specific reductions in cerebral blood flow (RCBF), indicating highly reproducible cortical hypoperfusion patterns.Regional Vulnerability Index (RVI):
Functional RVI (ReHo-derived) shows stronger and more consistent effects than any individual brain region and outperforms structural RVI measures.Large-scale cross-dataset validation:
Findings were replicated across 15,501 participants from UK Biobank, ENIGMA, Amish Connectome Project, and independent clinical cohorts, confirming robustness and generalizability.Biological interpretation via transcriptomics:
Integration of ReHo/RCBF with RNA-seq–based gene expression data reveals molecular mechanisms underlying functional deficits, highlighting synaptic, inflammatory, and neurovascular pathways.
🧬 Transcriptomics & Molecular Integration
- Brain-region–specific RNA-seq profiles mapped to functional deficits
- Differential expression of key synaptic, inflammatory, and neurovascular genes
- Pathway enrichment analyses (GO/KEGG) tied to neurofunctional alterations
- Integration of transcriptomic gradients with cortical maps for mechanistic insight
This positioning emphasizes transcriptomics as a central pillar, making the project highly relevant for researchers in multi-omics, systems biology, and computational psychiatry.
🔬 Functional Neuroimaging Biomarkers
- Regional Homogeneity (ReHo): voxel-level local synchrony in resting-state fMRI
- Regional Cerebral Blood Flow (RCBF): perfusion measures aligned with ReHo deficits
- Functional deficits in MDD consistently exceed structural cortical thickness changes
🔁 Workflow Schematic
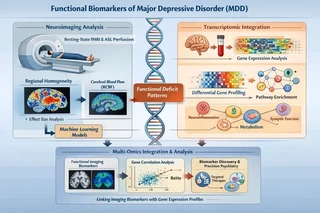
The schematic illustrates the multi-modal workflow:
- RNA-seq preprocessing and transcriptomic feature extraction
- Functional MRI preprocessing and ReHo/RCBF calculation
- Linking transcriptomic profiles with functional imaging deficits
- Integrating multi-modal features for biomarker discovery
- Mechanistic interpretation and predictive modeling
This framework provides biologically interpretable, cross-modal biomarkers suitable for diagnostics, stratification, and personalized interventions.
📌 Highlights (Summary)
- Functional deficits in MDD > Structural cortical reductions
- ReHo-based RVI shows strong reproducibility across datasets
- Transcriptomic integration links imaging phenotypes to molecular mechanisms
- Cross-validated over 15,501 participants
- Supports precision psychiatry approaches with multi-omics biomarkers
👥 Authorship & Credits
This project is based on a peer-reviewed research article published in JAMA Psychiatry. The present page emphasizes transcriptomics-informed modeling and computational extensions of the original findings.
Project Contributors:
Peter Kochunov, PhD
Professor of Psychiatry & Neuroimaging
Senior author, leading neuroimaging-genomics integration and transcriptomic interpretation.Syed Ibrar Hussain, PhD
Department of Mathematics, University of Houston, Houston, TX, USA Co-author of the original publication
Contributed to transcriptomics analysis, functional neuroimaging integration, and computational modeling.Additional Co-authors
Contributed to study design, data acquisition, transcriptomic profiling, neuroimaging analysis, and statistical modeling.
🤝 Collaboration & Contact
Open for collaboration in:
- Transcriptomics and RNA-seq integration
- Computational psychiatry and machine learning
- Multi-omics biomarker discovery
- Functional neuroimaging research
Contact via email or LinkedIn to discuss potential collaborations.
