βIV Spectrin & AKT/GSK3 Dysregulation as Multimodal Biomarkers in Schizophrenia
Project Overview
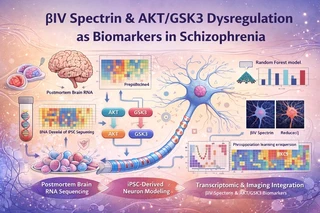
Schizophrenia (SCZ) is a severe neuropsychiatric disorder characterized by complex molecular, cellular, and functional brain abnormalities. Growing evidence indicates that functional and molecular dysregulation precede overt structural brain changes, motivating the development of biologically grounded, multimodal biomarkers.
This project integrates transcriptomics (RNA-seq), imaging genomics, and iPSC-derived neuronal models to characterize βIV spectrin (SPTBN4) and AKT/GSK3 signaling pathway dysregulation as convergent biomarkers of schizophrenia. By linking gene expression changes to cellular phenotypes and systems-level imaging features, the study provides a mechanistic and reproducible biomarker framework relevant for precision psychiatry.
🔗 Original peer-reviewed article:
βIV spectrin abundancy, cellular distribution and sensitivity to AKT/GSK3 regulation in schizophrenia – Molecular Psychiatry
🔑 Key Scientific Insights
- βIV spectrin reduction is a reproducible molecular signature of schizophrenia, observed in postmortem dorsolateral prefrontal cortex tissue and cellular models.
- AKT/GSK3 signaling directly regulates βIV spectrin stability, linking intracellular kinase dysregulation to neuronal cytoskeletal integrity.
- Transcriptomic co-expression analyses (PsychENCODE RNA-seq) reveal coordinated dysregulation of SPTBN4 and AKT/GSK3 pathway genes.
- iPSC-derived neurons from high-genetic-risk individuals recapitulate disease-associated molecular phenotypes, supporting causal relevance.
- Machine learning models integrating transcriptomic and imaging features achieve high classification accuracy (>95%), highlighting translational potential.
- Multi-omics convergence enhances biological interpretability, outperforming single-modality biomarkers.
🧬 Transcriptomics & Molecular Integration
Transcriptomics forms the biological foundation of this project:
- RNA-seq profiling of SPTBN4 (βIV spectrin) and AKT/GSK3 pathway genes
- Differential gene expression analysis in schizophrenia-relevant cortical regions
- Gene co-expression network analysis linking cytoskeletal regulation with intracellular signaling
- Pathway enrichment (GO / KEGG) implicating synaptic function, neuronal stability, and signal transduction
- Integration of transcriptomic gradients with imaging and cellular phenotypes
🔬 Functional, Cellular & Imaging Biomarkers
- βIV Spectrin (SPTBN4): Neuronal cytoskeletal protein critical for axonal stability and signal compartmentalization
- AKT/GSK3 Signaling Axis: Central intracellular pathway implicated in synaptic plasticity and psychiatric disorders
- iPSC-Derived Neurons: Disease-relevant cellular models enabling mechanistic validation
- Imaging Genomics: Links molecular dysregulation to systems-level brain phenotypes
- Machine Learning Models: Random Forest–based classifiers for multimodal biomarker prediction
🔁 Multimodal Workflow
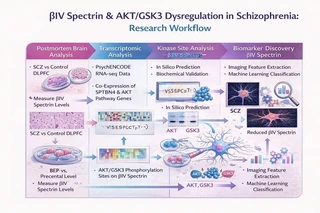
Pipeline overview:
- Postmortem brain RNA-seq and PsychENCODE transcriptomic analysis
- Identification of βIV spectrin and AKT/GSK3 pathway dysregulation
- iPSC-derived neuronal modeling and functional validation
- Imaging-genomics feature extraction and integration
- Machine learning–based biomarker classification
📊 Systems-Level Visualization
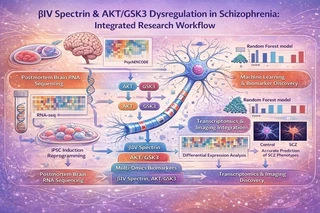
🧠 Clinical Relevance & Translational Impact
- Supports biologically interpretable biomarkers beyond symptom-based diagnosis
- Identifies cytoskeletal–signaling mechanisms as potential therapeutic targets
- Enables risk stratification using transcriptomics-informed features
- Provides a scalable framework for AI-driven precision psychiatry
🔮 Future Directions
- Expansion to single-cell and spatial transcriptomics
- Longitudinal imaging-genomics integration
- Cross-disorder analyses (e.g., bipolar disorder, MDD)
- Drug-response modeling targeting AKT/GSK3 signaling
- Translation toward clinically actionable biomarker panels
👥 Authorship & Credits
This project is based on a peer-reviewed publication in Molecular Psychiatry and is presented here as a computationally enhanced, transcriptomics-focused research extension.
Author Contributions
Fernanda Laezza, PhD
Professor of Pharmacology & Toxicology, University of Texas Medical Branch
Principal investigator and senior author; led experimental design, cellular modeling, and pathway analysis.Syed Ibrar Hussain, PhD
Department of Mathematics, University of Houston
Co-author of the original publication; contributed to transcriptomic integration, multimodal modeling, and computational interpretation.Additional Co-authors
Contributors to experimental neuroscience, transcriptomics, imaging analysis, and statistical modeling.
📄 Publication Reference
Laezza et al.
βIV spectrin abundancy, cellular distribution and sensitivity to AKT/GSK3 regulation in schizophrenia
Molecular Psychiatry (2025)
https://www.nature.com/articles/s41380-025-02917-1
🤝 Open to collaborations in transcriptomics, psychiatric genomics, iPSC modeling, imaging genetics, and AI-driven biomarker discovery.
